Lacuna - Correct Field Detection Challenge
Helping East Africa
$10 000 USD
Completed (almost 5 years ago)
Prediction
Earth Observation
634 joined
110 active
Start
Mar 26, 21
Close
Jul 04, 21
Reveal
Jul 04, 21
Data Updates
Data · 27 Apr 2021, 13:11 · edited ~3 hours later · 6
Hi Everyone
We just made the following updates to the competition dataset:
- Re-uploaded training and auxiliary data after removing duplicate data points which have the same image, plot size and corrected location.
- Added extra training data of 999 unique data points. You will find their images in separate folders so as not to re-download everything from scratch.
- Removed duplicate data points from the evaluation but kept them in test and sample submission files to keep everything in place on your side. We are currently re-evaluating all the submissions and will be done in a week.
- Added a separate folder with Sentinel-2 images for data points originally captured in 2015. These new images have 192 channels as the rest of the images.
- Modified the starter notebook to read and visualize the new data.
Great news!
But there is still the question on different markup (field location) for single image:
There are 4 labeled fields with the following IDs, but all of them lacated on the same image, we can build models that would be able to detect all of these 4 fields - it is not the problem.
But here we have 2 more unlabled IDs (fields) test=True in table above, and they are also lacated on the same image, and this is not so good, and makes possable solution not so good as can be expected, but anyway it is still solvable.
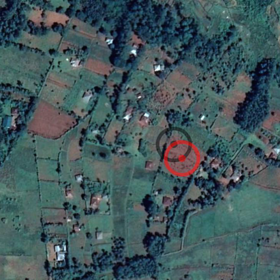
Real problems begins when we need to prepare predictions for fields located on the same image (here we have 2 IDs (fields), but there are samples with even more):
As you can see above for both test IDs we have same image and as a result we will have the same prediciton (predictions on the image just for demonstration purpose and does point to exact location of fields), and there is 2 ways (in case of 2 fields) of assigning fields to each ID - and its totaly random choise. And this can not be solved by droping duplicated, as in such case we still have several GT fields on the same image, and need to eandomly choose one for prediction.
This makes task unsolvable (at least for images where we have 2 or more fields located on the same image)
Hi AVoronov
Data samples that have the same images but different metadata like field size and yield are considered different since they represent different fields on the ground. I will make a post soon to the describe our annotation process to give you a better idea how important is the field size espicially to the problem.
Thanks for update.
Just to confirm (based on how you treat images with 112 bands in the new started notebook) - the *last* 16 bands represent the *most recent* month's spectral images?
Yes
Good news, one issue, starter Notebook is inassesible, it doesn't download
Press ctrl + s after clicking on the notebook link.