SUA Outsmarting Outbreaks Challenge
Helping Tanzania, United Republic of
$12 500 USD + AWS credits
Completed (over 1 year ago)
Prediction
810 joined
390 active
Start
Dec 06, 24
Close
Jan 31, 25
Reveal
Feb 01, 25
2nd Runners Up Approach
Notebooks · 6 Feb 2025, 08:50 · 5
Thanks @Amy_Bray and whole team @Zindi for everything you do. Congratulations also to all participants. The following is solo team Sinking Sand approach. Team name was adopted in anticipation of a major leaderboard shakeup that to some extent happened. Models not grounded on solid foundations would shake. I thought mine would be shaken badly and sink down the leaderboard-atleast now I see how lucky I was! Thanks to Zindi not including Cholera predictions in both public and private leaderboard evaluations.
INTRODUCTION
- All datasets are used
- No data reduction is done
- Only one model is used-Lightgbm
- No Post Processing is done to predictions
DATA CLEANING
- Duplicates: keep only the row with the maximum 'Total' value
- Drop nulls in water_sources-around 60 nulls
- Keep only the first four segments of facility ID to harmonize the column in both train and test
- Drop Lat and longitude features-did not have any predictive importance
MERGE
- Get anomalies & percent changes of all climate features in the extra datasets after grouping by 'lat_lon' before merging with train and test. These would later be powerful features for our models when we merge with train and test.
- Ckdetree was used to merge the data with the extra datasets
- Get 3 most nearest points to our location & also calculate their euclidian distances from our location.
- Integrate the climate features plus the anomaly and percent features of these three nearest points to our location
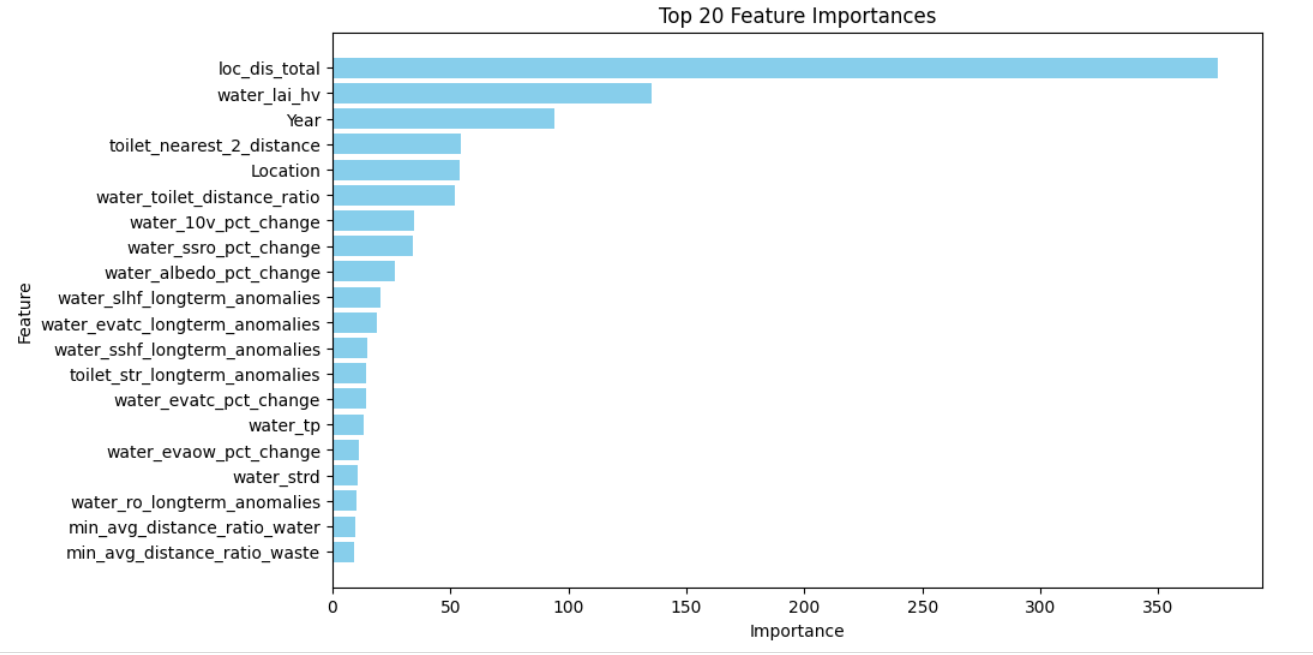
FEATURE ENGINEERING
- Monthly cyclical features-to capture the cyclical nature of months
- Used distances from the 3 nearest points from the location to create features such as Closest distance to water, toilet, and waste facilities of a certain location
- Group target encodings e.g. train.groupby(["Location", "Disease"])["Total"].median(). The problematic location (ID_0358ea0e-af2d-450e-8124-4144ca7860fa) in test but not in train was left as is (with NaNs)
- Geographical clustering using kmeans.
MODELLING
- Many models were tried initially, Lightgbm Regressor worked best comparatively
- Simple Train Test Splitting worked best comparitively
- Log transforming target was better than having raw targets
- A different fitting is done for different diseases
- On cross-validation:Diarrhoea had the highest MAE while dysentry had the least
- On test: Public Lb-around 5.6, Private LB-around 6.6
- No post processing was done
FINAL THOUGHTS
- Location & Disease grouped target encoding was the best feature in all disease scenarios
- I still feel data reduction if explored could have improved the results further
- Tuning lgbm parameters could improve the results further. Was not able to use provided credits due to errors-did not send an email for help though, my mistake!
It's loaded XD. Good job and congratulations!
Congratulations and thanks for sharing Julius!
Thanks for sharing and congratulations!
thanks for sharing and congrats
Thank you good people. You all did very well too brothers.